
IP-003 based on Fluorescence Spectrography Method
Fluorescence spectrography, also called fluorescence spectroscopy, is an analytical technique used to measure the fluorescence emitted by a substance when it absorbs light. It is commonly used to study molecules such as proteins, DNA, and other biologically relevant substances. Principle:
Excitation: The sample is first exposed to light of a specific wavelength (excitation wavelength). This energy excites the electrons in the molecules to a higher energy state.
Emission: After absorbing the excitation light, the molecules will relax and emit light (fluorescence) at a longer wavelength than the excitation light.
Detection: The emitted fluorescence is detected using a spectrograph or a photomultiplier tube, and its intensity is measured across various wavelengths.
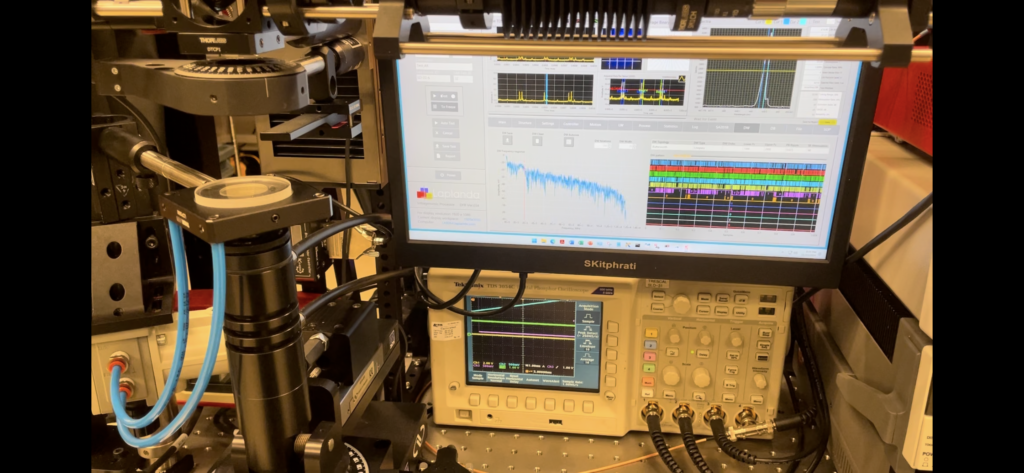
Steps in Fluorescence Spectroscopy:
- Sample Preparation: The substance to be analyzed is either naturally fluorescent or tagged with a fluorescent marker (fluorophore).
- Excitation Light Source: A light source, such as a laser or xenon lamp, is directed at the sample to excite the molecules.
- Emission Detection: The emitted light is captured by detectors, and a spectrum is generated showing the fluorescence intensity versus wavelength.
- Analysis: The emitted fluorescence spectrum provides information about the molecular structure, environmental changes, or concentration of the target substance.
Fluorescence spectroscopy is widely used in biological research, environmental monitoring, and clinical diagnostics due to its high sensitivity and specificity.
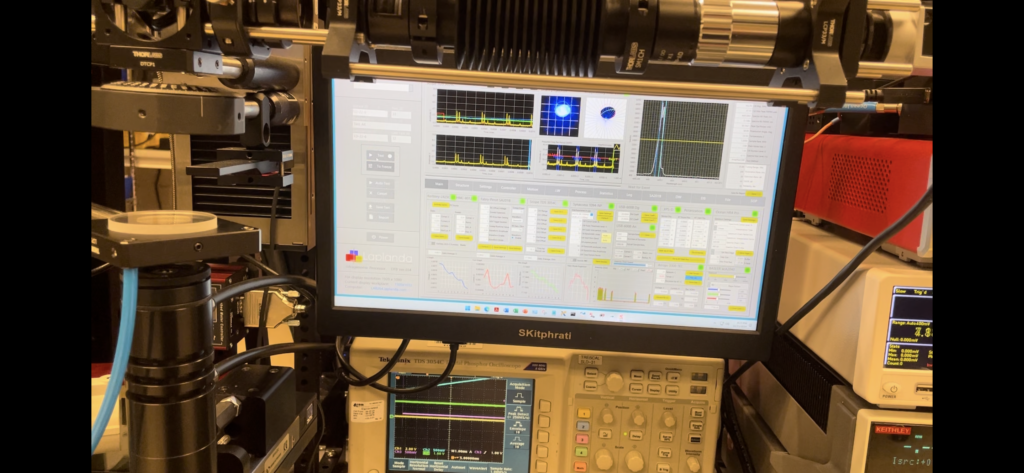

Fluorescence-Based Sequencing (e.g., Sanger Sequencing and Next-Generation Sequencing)
Fluorescence-based sequencing refers to the use of fluorescence to identify the sequence of nucleotides (A, T, C, G) in DNA. One of the most common methods that utilize fluorescence for sequencing is Sanger sequencing and Next-Generation Sequencing (NGS) technologies.
1. Sanger Sequencing (Chain Termination Method)
Sanger sequencing is a method for determining the sequence of nucleotides in DNA. It uses fluorescently labeled dideoxynucleotides (ddNTPs) to terminate the DNA elongation at specific points.
Steps in Sanger Sequencing:
- DNA Template: The DNA fragment of interest is used as a template.
- Primer Binding: A short primer binds to the template DNA to initiate replication.
- DNA Synthesis: DNA polymerase extends the primer, synthesizing a complementary strand. However, the synthesis can be terminated if a dideoxynucleotide (ddNTP) is incorporated instead of a regular nucleotide (dNTP).
- Fluorescently Labeled ddNTPs: The ddNTPs are labeled with different fluorescent dyes for each of the four nucleotides (A, T, C, G). Each dye emits light at a different wavelength when excited by a laser.
- Separation and Detection: The products of the DNA synthesis are separated by size using capillary electrophoresis. The DNA fragments are passed through a detector, which reads the fluorescence signal to determine the nucleotide sequence.
This process produces a chromatogram, which can be analyzed to determine the nucleotide sequence of the DNA.
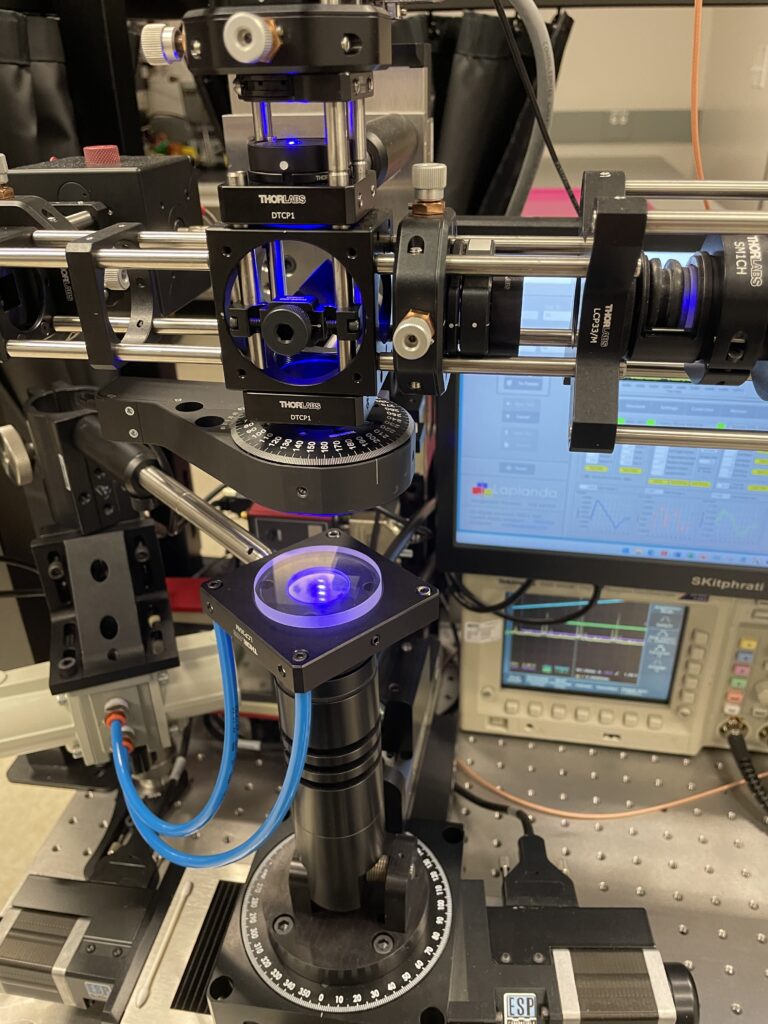

2. Next-Generation Sequencing (NGS)
Next-Generation Sequencing is a more advanced and high-throughput technology for sequencing large amounts of DNA quickly and at a lower cost compared to Sanger sequencing. NGS uses fluorescently labeled nucleotides to detect DNA synthesis in real-time.
Steps in NGS:
- DNA Fragmentation: The DNA is fragmented into smaller pieces.
- Adapter Ligations: Specific adapters are ligated to the ends of the fragments to allow binding to a sequencing flow cell.
- Amplification: The fragments are amplified to create clusters of identical DNA sequences.
- Sequencing: During sequencing, the DNA fragments are extended one base at a time. Each nucleotide (A, T, C, G) is labeled with a different fluorescent dye, and when a nucleotide is incorporated into the growing strand, the corresponding fluorescence is detected.
- Data Analysis: The fluorescence signals are captured and analyzed to determine the DNA sequence. This results in high-throughput data that can be assembled and analyzed.
NGS is widely used for genome sequencing, transcriptomics, and other applications that require large-scale DNA sequencing.
Applications of Fluorescence Spectroscopy and Sequencing:
- Fluorescence Spectroscopy: Used in protein quantification, detection of pollutants, cellular imaging, and monitoring biochemical reactions.
- Fluorescence Sequencing: Used for DNA sequencing, mutation detection, genotyping, and transcriptome analysis.
Both techniques are fundamental in modern molecular biology, providing powerful tools for analyzing and understanding biological systems.